BHQplus™ Probes are compact, dual-labelled, hydrolysis probes for qPCR. This probe type features a 5' fluorescent dye and a 3' Black Hole Quencher™ (BHQ) with the unique BHQplus duplex stabilising chemistry to enhance specificity and mismatch discrimination.
BHQplus Probes use modified C and T nucleotides to stabilise the probe-target duplex, which enables design of shorter oligonucleotides, typically 15 to 25 bases.
Key benefits:
- Short probes with high specificity: Duplex stabilisers permit the design of shorter dual-labelled probes with increased stability and specificity during hybridization.
- Ideal for difficult targets: High duplex stability improves SNP/mismatch discrimination and binding to difficult regions including AT-rich targets.
- Improved assay sensitivity: Our best-in-class BHQ is positioned closer to the dye, resulting in greater quenching efficiency.
- Versatile: These probes can be paired with a wide range of dyes to fit your exact instrument and application.
Greater duplex stability for enhanced specificity and sensitivity
BHQplus Probes form more stable probe-target duplexes, enabling shorter oligonucleotide designs. Shorter oligos offer high specificity and better detection of difficult targets, such as AT-rich sequences. A shorter BHQplus probe will generate lower background and exhibit a higher signal-to-noise ratio, enhancing the sensitivity of your assay.
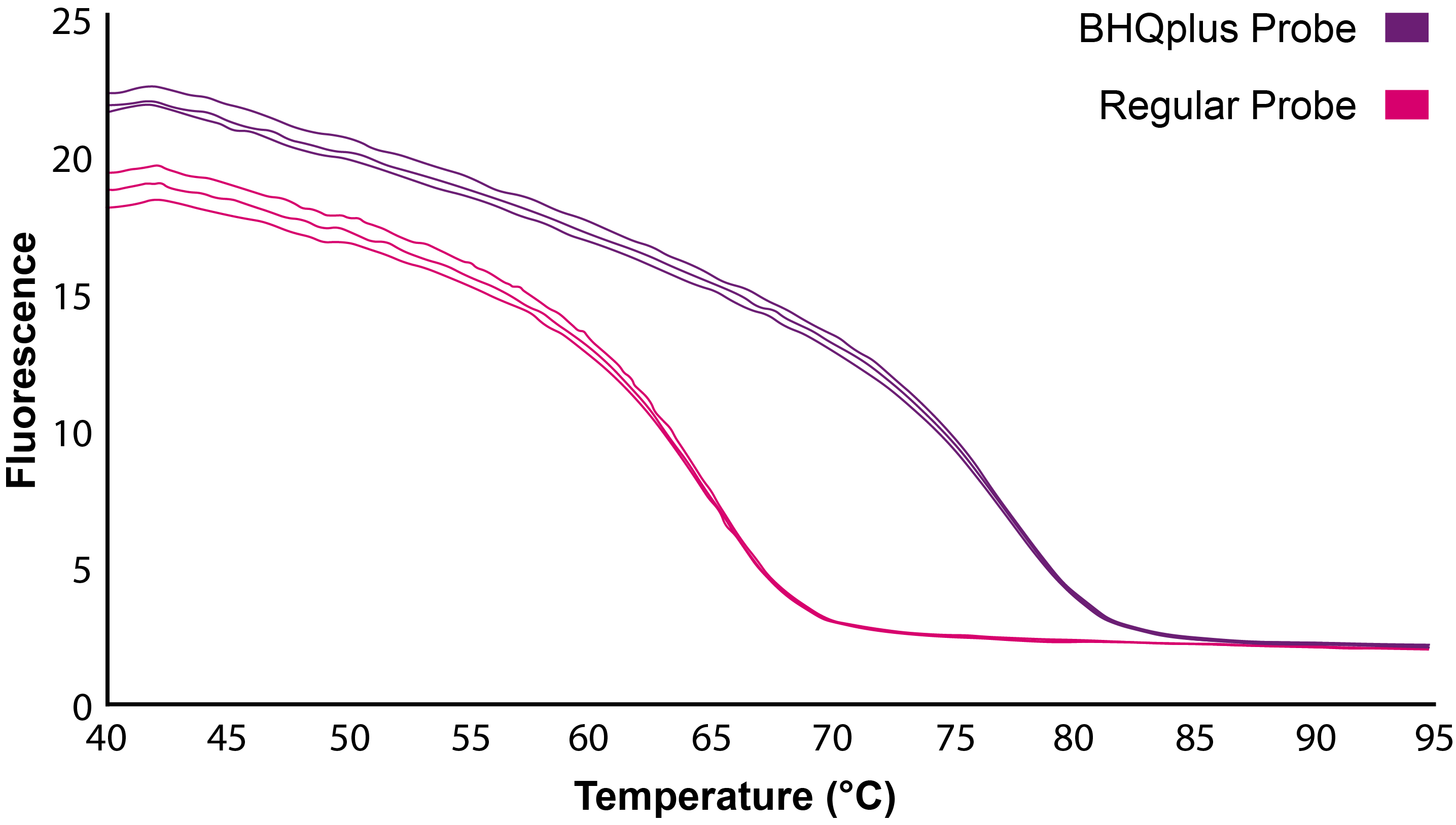
This melting temperature graph shows an example of a BHQplus Probe denaturing at a temperature >10 °C higher than the denaturation of a regular probe, indicating the increased stability of the BHQplus-target duplex.
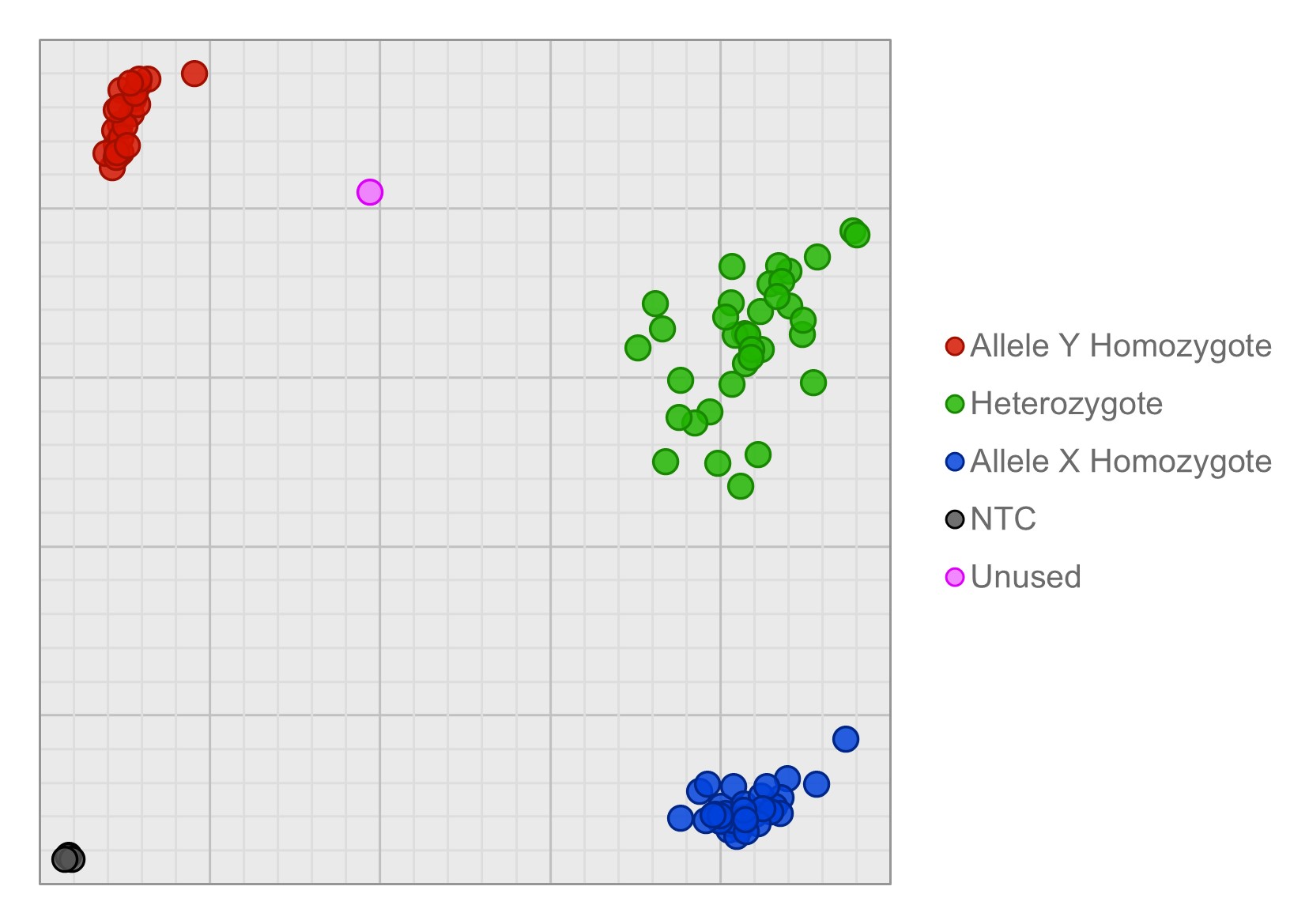
SNP genotyping cluster plot illustrating HotShot extracted maize DNA run with BHQ Probe Master Mix, with maize-specific BHQplus Probe-based assays. The BHQplus Probes were labelled with either FAM (specific for allele X) or CAL Fluor Orange 560 (specific for allele Y). Each data point represents a 1 µL total reaction volume in a 1536-well plate, with the experimental protocol run on the SNPline instrument platform. This experiment returned a 98.9% call rate, out of 96 total samples tested. This demonstrates high specificity and sensitivity of BHQplus Probes, generating high quality data from crude-extracted DNA on a complex plant species.
Achieve optimal quenching efficiency
When it comes to assay design, we don’t believe in a one-dye-fits-all strategy. Our wide selection of dyes empowers you to design the best PCR or qPCR probe for your application without sacrificing performance. We have paired each dye with the BHQ quencher that offers optimal quenching efficiency.
|
5' fluorescent dye |
Abs (nm) |
Em (nm) |
3' quencher |
|
|---|---|---|---|---|
| FAM | 495 | 520 | BHQ-1 | |
| TET | 521 | 536 | BHQ-1 | |
| CAL Fluor Gold 540 | 522 | 544 | BHQ-1 | |
| CIV-550 | 530 | 550 | BHQ-1*, BHQ-2 | |
| HEX | 535 | 556 | BHQ-1 | |
| CAL Fluor Orange 560 | 538 | 559 | BHQ-1 | |
| CAL Fluor Red 610 | 590 | 610 | BHQ-2 | |
| Quasar 670 | 647 | 670 | BHQ-2 | |
* Recommended quencher
Use the spectral overlay tool to view compatible dyes for your instrument. Learn more
 Order our premixed ValuMix assays to streamline your experimental setup while reducing pipetting errors
Order our premixed ValuMix assays to streamline your experimental setup while reducing pipetting errors
ValuMix assays for qPCR and gene expression
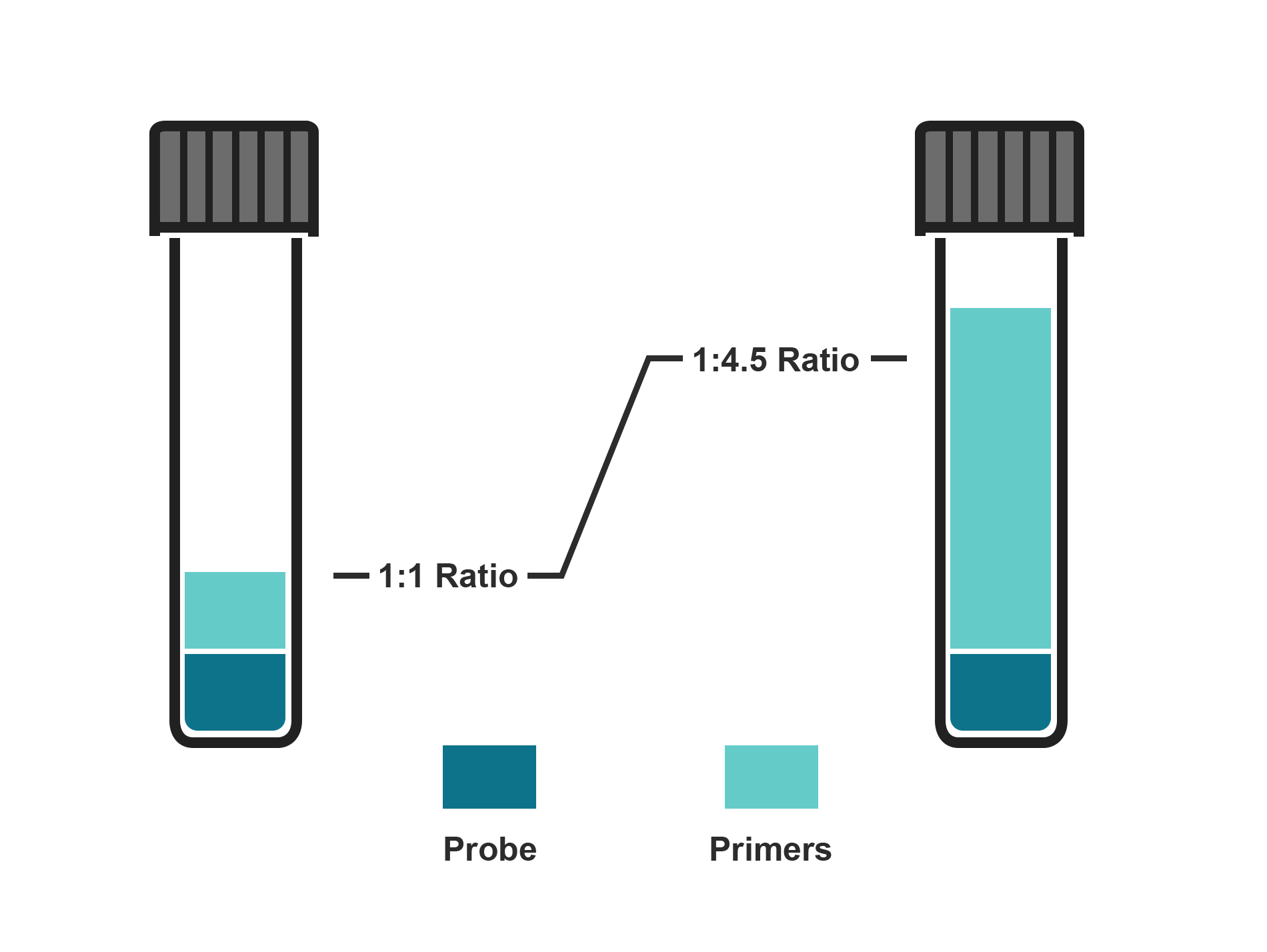
This format contains one custom BHQplus Probe and two primers combined in a single tube. Adjust primer quantities based on probe amount and select a probe:primer ratio between 1:1 and 1:4.5.
Key features
- Probes purified via RP HPLC
- Salt-free primers
- Wide selection of dyes: FAM, TET, CAL Fluor Gold 540, CIV-550, HEX, CAL Fluor Orange 560, CAL Fluor Red 610 and Quasar 670
- Probe quantities: 0.5 nmol (FAM only), 5 nmol, or 20 nmol
ValuMix assays for SNP genotyping
This ValuMix assay format contains two custom BHQplus Probes and two primers combined in a single tube, with a probe to primer ratio of 1:4.5.
Key features
- Probes purified via RP HPLC
- Primers purified via RPC
- Probe 1: FAM
- Probe 2: Choose from TET, CAL Fluor Gold 540, CIV-550, HEX or CAL Fluor Orange 560
- Size options: small (2 nmol/probe, 9 nmol/primer), medium (5 nmol/probe, 22.5 nmol/primer) and large (12 nmol/probe, 54 nmol/primer)
- Available as a 40X solution at 8 μM/probe and 36 μM/primer
Design short BHQplus Probes with ease
Are you designing a new assay or considering BHQplus probes for a sequence currently designed for a different probe technology? For design assistance to achieve optimal results, please contact our assay design service team to redesign your assay with BHQplus probes.
Redesigning your assay for a BHQplus probe is required if you are using a traditional probe sequence that is longer than 25 bases or that has a predicted melting temperature of 70 °C. Alternatively, consider ordering the sequence as a BHQ Probe (traditional length) or BHQnova Probe (extended length).
For design or technical assistance, please contact our assay design and support team with questions regarding probe design.